OASEGSAR
Segment an image
Description
OASEGSAR applies a hierarchical region-growing segmentation to SAR image data and writes the resulting objects to a vector layer. Each object represents a homogeneous local area according to the input channels and the definition of criteria of homogeneity based on the scale, shape and compactness.
Parameters
oasegsar(fili, dbic, segeigen, sarfilt, flsz, trimrt, maskfile, mask, filo, ftype, dbsd, segscale, segshape, segcomp)
| Name |
Type |
Caption |
Length |
Value range |
| FILI* |
str |
Input file name |
1 - |
|
| DBIC |
List[int] |
List of input channels |
0 - |
|
| SEGEIGEN |
str |
Eigenvalues for segmentation |
0 - |
L1 | L2 | L3 |
| SARFILT |
str |
SAR filter type |
0 - |
ELEE | Boxcar | None
Default: ELEE |
| FLSZ |
List[int] |
Filter size |
0 - 1 |
5 | 7 | 9 | 11 | 13 | 15 | 17 | 19 | 21 | 23 | 25 | 27 | 29 | 31 | 33
Default: 5 |
| TRIMRT |
List[int] |
Percentage to trim right tail |
0 - 1 |
0 - 50
Default: 15 |
| MASKFILE |
str |
Name of input AOI file |
0 - |
|
| MASK |
List[int] |
Layer number of AOI |
0 - 1 |
|
| FILO |
str |
Output file name |
0 - |
|
| FTYPE |
str |
Output file type |
0 - 3 |
PIX | SHP
Default: PIX |
| DBSD |
str |
Output segment description
|
0 - 64 |
Default: Segmented Layer |
| SEGSCALE |
List[float] |
Scale of objects |
0 - 1 |
5 - 1000 |
| SEGSHAPE |
List[float] |
Shape of objects |
0 - 1 |
0.1 - 0.9 |
| SEGCOMP |
List[float] |
Compactness of objects |
0 - 1 |
0.1 - 0.9 |
* Required parameter
Parameter descriptions
FILI
The name of a GDB-supported file that contains the image channels to use for segmentation.
DBIC
A list of channels in the raster image to process.
By default, all channels are processed.
SEGEIGEN
The image-covariance eigenvalues to use in the image-segmentation process.
You can use all or a subset of the following image-covariance eigenvalues:
-
L1 (\u00ce\u00bb1)
-
L2 (\u00ce\u00bb2)
-
L3 (\u00ce\u00bb3)
To use the eigenvalues, the input SAR image must be single-look complex (SLC) and fully polarimetric.
SARFILT
The SAR filter to use to filter the input channels prior to segmentation.
You can specify either of the following filters:
-
ELEE (Enhanced Lee)
-
Boxcar
For information about the filter types, see Data filtering and tail trimming.
If you have already run filtering on the input SAR file, you need not specify a value for this parameter.
FLSZ
The size of the filter window, in pixels and lines, that is moved across the input image to determine the output pixel value.
The value you specify must be an odd integer and a minimum of 5. Increasing the value increases the processing time at an approximate quadratic rate.
TRIMRT
The percentage to trim the right tail of the data distribution.
The default value is 15.
MASKFILE
The file that has the vector layer containing the area of interest (AOI) to process.
This parameter is optional.
MASK
The number of the vector layer that contains the AOI to process.
A vector layer can contain multiple AOIs. Each must be a closed polygon.
If you specify a value for MASKFILE, you must specify a value for this parameter.
FILO
The name of the output file to which to write the segmentation.
If you do not specify a value, a new vector later containing the segmentation is written to the input file you specified.
FTYPE
The format of the output file.
The following formats are supported:
The default is PIX.
DBSD
Describes (in up to 64 characters) the contents or
origins of the output layer.
SEGSCALE
The scale parameter of the multiresolution segmentation algorithm.
Scale is a unitless parameter that controls the average object size during the segmentation process. A small scale results in a greater number of objects at the expense of the generalization. A large scale forces the segmentation to create fewer objects, but various types of land cover, for example, might be merged together.
SEGSHAPE
The shape parameter of the multiresolution segmentation algorithm.
A low shape value, such as 0.1, places a high emphasis on color; that is, pixel intensity, which is typically the most important aspect of creating meaningful objects.
SEGCOMP
The compactness parameter of the multiresolution segmentation algorithm.
A high compactness weighting, such as 0.9, produces more compact object boundaries, such as with crop fields or buildings.
Details
OASEGSAR uses an open-source segmentation method, known as a multiresolution segmentation algorithm.
The first step in object-based image classification is segmentation: calculating discrete regions of image objects. This is achieved by stratification of an image. These image-objects are used further as the basic unit of analysis for developing image-analysis strategies, including classification and change detection.
To date, various image-segmentation techniques have been developed with mixed results. The earlier developments in segmentation techniques, however, were in the machine-vision domain and aimed at the analysis of patterns, the delineation of discontinuities on materials or artificial surfaces, and the quality control of products. Later, these concepts were used in remote sensing (RS) for attribute identification.
Functionally, the segmentation process partitions an image into statistically homogenous regions or objects (segments) that are more uniform within themselves and differ from their adjacent neighbors. Another conceptually important aspect of image segmentation is its relevance to spatial-scale theory in RS, which describes how local variance of image data in relation to the spatial resolution can be used to select the appropriate scale for mapping individual land-use classes. Image segmentation defines the size and the shape of image objects and influences the quality of the follow-on analysis: classification.
However, a single segmentation algorithm does not yet exist that can reproduce the same objects as a human can identify intuitively. To attempt to overcome this issue, image objects are created that are as homogeneous as possible and grouped together by using a classification process to create objects that can come close to human perception. This can be an iterative process in which various segmentation algorithms are used, or a specific algorithm is used with various parameters, to achieve a suitable result.
A data set of input imagery contains multiple channels. The underlying algorithm is a hierarchical step-wise region grown by using random "seeds" spread over the entire image. This method can be classified as a "bottom-up" optimization routine that starts with a pixel and ends with segments that are groups of like pixels.
The criteria that defines the growth of a region can be based on the difference between the pixel intensity and the mean of the region. The algorithm assesses the local homogeneity based on the spectral and spatial characteristics. The size of the object is controlled with the value for scale, which you specify. The larger the scale, the larger will be the output object. Other homogeneity criteria are based on shape and compactness.
The result of the segmentation process is as follows:
- An initial abstraction of the original data
- Creation of a vector (polygon) representation of the image objects
The polygons, at this level of abstraction, can be considered image objects in the image domain and do not necessarily represent objects in the real world.
Required available memory
Segmentation is intense computationally and, as such, relies on the amount of memory (RAM) on your computer. The amount of memory used depends on the size (in pixels and channels) of the input raster. To calculate the amount of RAM (in gigabytes) that will be required by
Object Analyst to process a file, you can use the following formula:
- nGB = nX × nY × (4 × nC + 29) ÷ 795364314.1
- where:
- nGB is the amount of required memory (RAM in gigabytes) to perform the segmentation
- nX is the number of pixels in the x-dimension (pixels)
- nY is the number of pixels in the y-dimension (lines)
- nC is the number of channels
Note: The segmentation algorithm respects NoData pixels in the source image. That is, a pixel defined as NoData by the NO_DATA_VALUE metadata tag will be excluded from the segmentation process. The file-level metadata tag is used as the default for each channel; however, channel-level tags, when available, will override the default. When metadata does not exist, each pixel in the input is considered valid.
SAR-imagery suppport
You can process SAR data in its raw vendor format or, alternatively, you can ingest it to a PCIDSK (.pix) file for processing.
SAR imagery comes in many formats and processing levels. The specific SAR-data type and its processing level are identifiable by the matrix type in the image metadata. The matrix type determines which, if any, SAR preprocessing parameters you can apply, and which SAR attributes you can calculate, as shown in the following table.
Table 1. SAR matrix and channel types, segmentation processing, and predefined attribute types to calculate
| Matrix type |
Channel type** |
Segmentation, preprocessing |
Attribute types to calculate |
| [c1r] Single polarization, detected |
1 x [16U] (uncalibrated)
- or -
1 x [32R] (calibrated)
|
• Filtering
• Tail trimming
|
• Statistical
• Geometrical
• Texture
|
| [c1c] Single polarization, complex |
1 x [C16S] (uncalibrated)
- or -
1 x [C32R] (calibrated)
|
• Filtering
• Tail trimming
|
• Statistical
• Geometrical
• Texture
|
| [c2r] Dual polarization, detected |
2 x [16U] (uncalibrated)
- or -
2 x [32R] (calibrated)
|
• Filtering
• Tail trimming
|
• Statistical
• Geometrical
• Texture
|
| [s2c] Dual polarization, complex* |
2 x [C16S] (uncalibrated)
- or -
2 x [C32R] (calibrated)
|
• Filtering
• Tail trimming
|
• Statistical
• Geometrical
• Texture
|
| [s4c] Fully polarimetric, complex |
4 x [C16S] (uncalibrated)
- or -
4 x [C32R] (calibrated)
|
• Filtering
• Tail trimming
• Extra segmentation layers:
- L1 (λ1)
- L2 (λ2)
- L3 (λ3)
|
• Statistical
• Geometrical
• Texture
• Polarimetric
|
* Compact-polarimetric data is a special case of [s2c] data.
** Calibrated data is recommended.
With a given image, you can run segmentation on all channels or a subset thereof.
Note: Preprocessed or modified SAR images containing a mix of detected and complex channels cannot be segmented into objects.
With fully polarimetric complex data [s4c], the following additional layers are available for segmentation:
The preceding layers correspond to the three eigenvalues of the covariance matrix of each image and can enhance land-cover attributes that are not always distinguishable from the original HH, HV, VH, and VV channels.
Data filtering and tail trimming
Enhanced Lee applies a Lee adaptive-speckle filter. While reducing speckle in the image, the filter preserves the polarimetric and spatial information in the image. Spatial information is preserved by using an edge detector to identify a homogeneous local neighborhood in which to estimate the filter parameters.
Boxcar applies a boxcar filter to detected or complex-valued SAR data. It is used commonly to increase the effective number of looks (ENL) in single-look or multi-look SAR data. Speckle reduction is applied by using local averaging.
Note: It is recommended that you filter your SAR data before running segmentation on it.
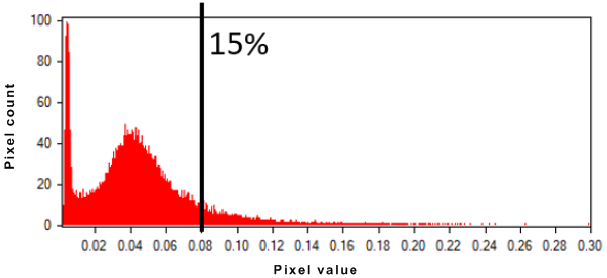
To minimize the effect of backscattering, you can apply tail trimming. With SAR data, frequency histograms are often characterized by strong outliers in the rightmost tail of the data distribution; that is, point targets with a high-backscattering coefficient several magnitudes higher than the bulk of the data. Segmentation is influenced strongly by these "scatterers" and can produce substandard results over distributed targets, such as forest, water, agricultural field, bare soil, and others.
The following image, for a given SAR channel, shows that 15 percent of the brightest pixels correspond to 0.08 in intensity. With at tail trim of 15 percent, all pixels with a value higher than 0.08 will be reassigned temporarily to 0.08 for the segmentation; however, the original pixel values are used in the attribute calculation.
Figure 1. Tail trim
Examples
In the following example, all polarimetric channels (1: HH, 2: HV, 3: VH, 4: VV) and the eigenvalues ("L1, L2, L3") of a fully polarimetric Radarsat-2 image are used for segmentation. An Enhanced Lee filter (ELEE), with a 9 x 9 window, and a tail trim of 15 percent are applied to all channels prior to segmentation.
from pci.oasegsar import oasegsar
fili="RS2_SLC_FQ09subset.pix"
dbic= [1,2,3,4]
segeigen= "L1, L2, L3"
sarfilt= "ELEE"
flsz= [9]
trimrt= [15]
maskfile= ""
mask= []
filo= "RS2_SLC_FQ09subset_seg_35_0.1_0.5.pix "
ftype= ""
dbsd = ""
segscale= [35]
segshape= [0.1]
segcomp= [0.5]
oasegsar (fili, dbic, segeigen, sarfilt, flsz, trimrt, maskfile, mask, filo, ftype, dbsd, segscale, segshape, segcomp)
In the following example, the VV and VH detected channels of a Sentinel-1 image, in ground range, are used for segmentation. Am Enhanced Lee filter (ELEE), with an 11 x 11 window, and a tail trim of 15 percent are applied to all channels prior to segmentation. An AOI (maskfile) is used to constraint the segmentation to a specific area of the input Sentinel-1 image.
from pci.oasegsar import oasegsar
fili="S1A_IW_GRDH_VV_VH_subset.pix"
dbic= [1,2]
segeigen=""
sarfilt= "ELEE"
flsz= [11]
trimrt= [15]
maskfile= "S1A_IW_GRDH_VV_VH_subset_OA_Area_of_Interest.pix"
mask= [2]
filo="S1A_IW_GRDH_VV_VH_subset_seg15_0.1_0.5.pix"
ftype= ""
dbsd=""
segscale= [15]
segshape= [0.1]
segcomp= [0.5]
oasegsar (fili, dbic, segeigen, sarfilt, flsz, trimrt, maskfile, mask, filo, ftype, dbsd, segscale, segshape, segcomp)
References
For more information about region-growing segmentation, scale, shape, and compactness parameters, see the following published works:
- Baatz, Martin, and Arno Schäpe. "Multiresolution segmentation: an optimization approach for high quality multi-scale image segmentation." Angewandte Geographische Informationsverarbeitung XII 58 (2000): 12-23.
- Dey, Vivek, Yun Zhang, and Ming Zhong. A review on image segmentation techniques with remote sensing perspective. na, 2010.
© PCI Geomatics Enterprises, Inc.®,
2024. All rights reserved.
